Documented on this page is a Princeton setup for improving the quality of microarrays by ozone remediation using a catalyst filter. I'm hoping other people can benefit from my experiences. If you have any additions to the annals of microarray misery, or any corrections to what I've posted, please email me at maitreya [at] princeton [dot] edu.
FYI, my latest microarray protocols.
Fall-winter 2003-2004, right after I moved to Princeton from Stanford, I was getting very nice arrays. As 2004 progressed, my arrays started getting worse and worse. In particular, my Cy5 signal decreased precipitously. Since the same protocols had worked so well just months before, I finally concluded that it must be seasonal. That narrowed it down to temperature, humidity, and ozone.
Agilent was the first to explicitly test the effects of ozone on cyanine dyes (in particular, the Amersham Cy dyes and the Molecular Probes Alexa dyes). You should read their paper for more details. (Email me if you can't find Analytical Chemistry back issues, and I'll send you the PDF.) The take-home message of the paper was that 5 ppb or above of ozone for more than 5 minutes will annihilate Cy5. They also have a really nice Technical Note on the effects and how to combat them (also in a shorter version), and an e-seminar.
I've been told by Dan Hogan at Stanford that he's also tested Cy5Star and 650 Oyster with similarly bad results. Dan's results also show some pretty scary spurious clustering based on ozone levels.
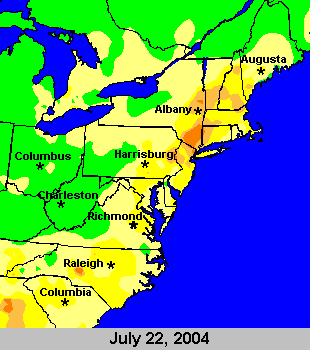
You can find out the ballpark local ozone levels from various sources on the web. My favorite for Princeton is the NJDEP Bureau of Air Monitoring since it has actual numbers in addition to the health levels. Conviently, the AQI is almost exactly ppb for ozone. There are also fun maps from the EPA that show ozone. Here's a nice example of the peak levels on July 22:

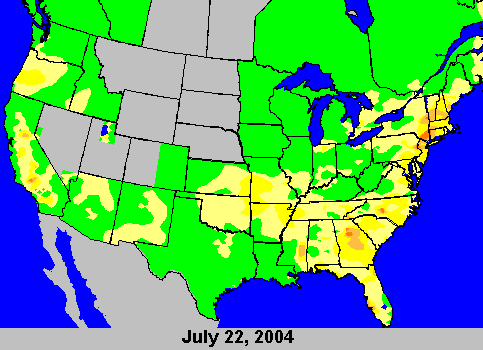
Note that to get above green on the map, you have to be over 60 ppb. The darkest orange is 111-124 ppb. By then, your Cy5 is long gone.
Weather Underground maps are also nice because they show gradations of color in about 10 ppb intervals.
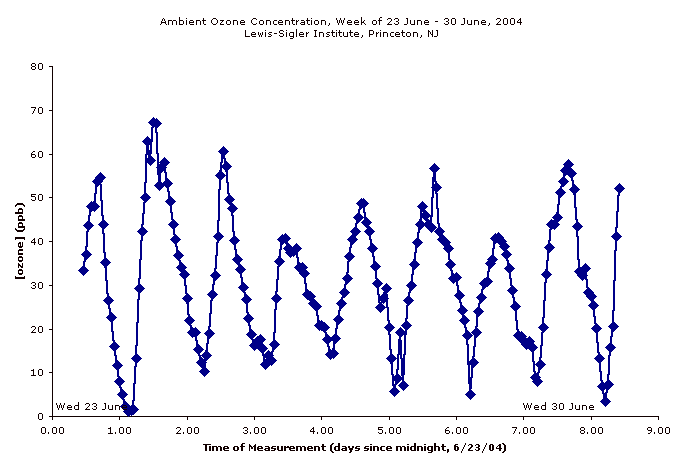
The core facility at Princeton has a Teledyne 400E ozone detector, so we set it up to read the levels at my bench. Around 2:00 in the afternoon, right about the time I generally do my array work, the levels were ~60 ppb. I concluded the problem was probably ozone and decided to make an ozone-free room.
Jeff Cross at Tri-dim filters (856-786-2447) recommended a carbon filter, which is also what Agilent concluded. I bought one of their 300-500 CFM units for about $2000 and hooked it up in a room with a door. The levels of ozone dropped from about 60 ppb to about 10. We had to block off the air conditioning vents to get down the last 10 ppb. Opening and closing the door occasionally doesn't seem to raise the levels. (Note: this filter unit puts out A LOT of dust. Make sure you get it with an additional particulate filter if you go this route. Keep reading for the solution we recommend.)
I did 2 arrays to test the setup. At my bench I used RT to make amino-allyl cDNA. I split the cDNA for 2 self-self arrays. I resuspended 1 tube of Cy3 and 1 tube of Cy5 in the ozone free room. I added an aliquot to each cDNA aliquot. Then I carried the rest of the dye and cDNA to my bench and set up a second set of labeling reactions. All the steps were continued in parallel until the scan, which was all done in the ozone free room.
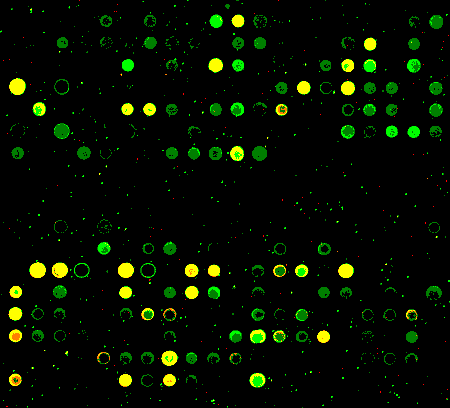
at my bench
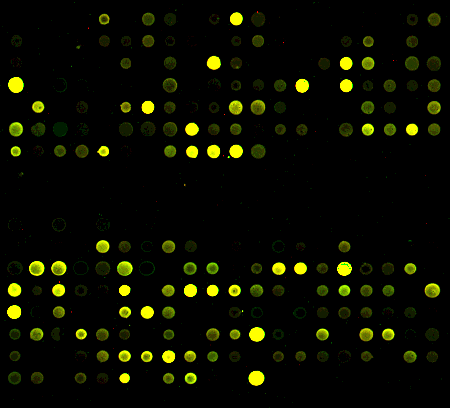

with a carbon filter
These are scatterplots of the Cy3 vs Cy5 net intensity for the 2 experiments. Note that I've maxed out the red channel on the carbon filter array.
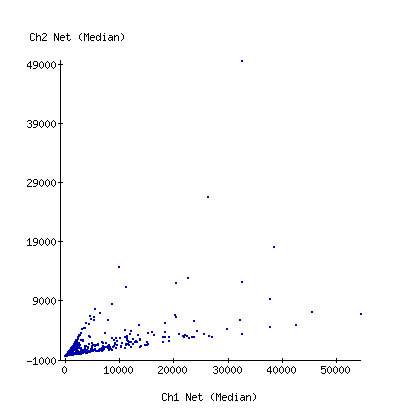
at my bench
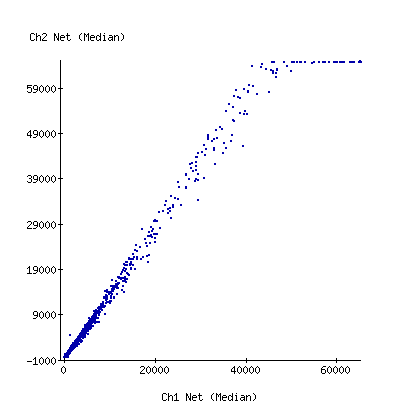
with a carbon filter
Before we found the catalyst filters (see below), we improved the carbon filtration setup for a larger room. (I'm leaving these materials in just in case people need alternatives or additions to the catalyst.) We added 3 woven carbon filters, one on the ceiling air vent (air circulation is turned down, too), and two taped to box fans. One of the box fans was a $10 KMart fan, and the other was a couple hundred dollar heavy duty fan. You could tell that the wimpy fan didn't push as much air through the filter as the heavy duty one (and it died after about 3 months of constant use). The woven filters were Tri-dim 24x24x2 Tri-Sorb XL Heavy Duty Pleated Carbon Filters, part 69224242H. They're cheap, like $6 each or something. The 2 inch thick woven filters by themselves weren't sufficient to keep the ozone low. There are also thicker carbon filters that may be better. The filters need occasional replacement, like every 3 months or even longer.
Amersham seems to still deny that ozone is a problem. (See this Application note.) All I know is that my problems are seasonal and solved by a carbon filter.
Since the initial experiment, we've improved the setup to make the entire microarray core facility ozone-free. Donna Storton in the core facility has posted a schematic of our current filtration ductwork. The big breakthrough was finding an ozone catalyst filter, which works way better than carbon. The air conditioning is back on, for example. BASF Catalyst LLC (formerly Engelhard Corp.) makes the catalyst. Talk to Vince Patram (732-205-6236), not the person listed on the schematic.
Pat Brown's lab at Stanford has a similar solution, which they call the ozone annihilator. Their specs are to build a unit to encase a scanner. Bruce Futcher's lab in New York and the Johns Hopkins microarray facility both purge their scanners with nitrogen for a similar result. We went with the ozone-free room solution because even the benchtop work is sensitive to the extreme New Jersey ozone levels. My friend Jason Lieb has also been able to implement an ozone-free room at UNC Chapel Hill, using box fans and the catalyst filters.
Companies are also starting to catch on. There are a few different systems available. I'm not linking them because they are pretty sketchy and I have not heard any good recommendations of them. Invitrogen seems to be trying to find ozone-resistant dyes.
If you can't make an ozone-free space, there are a few reagents around that claim to help. One is from Agilent (Stabilization and Drying Solution, part number 5185-5979) and the other is from Genisphere (Dyesaver, part Q500500). Amersham includes a homemade recipe in this Application note. All involve dipping your hybed and washed array in the substance. I think they may be solutions where ozone levels are not quite as high as they are here, or for people who can't convince the owner of a shared scanner to make it ozone-free. I have not personally tried any of them, but I have gotten several first hand accounts of good results with the Agilent reagent. According to Dan Hogan at Stanford, it tends to make homemade arrays look a little swirly, but the increased data quality makes up for it.
I hate Cy5. You can get your very own I hate Cy5 merchandise at my Cafe Press shop. I don't make any money off this, by the way. I just hate Cy5 and I bet you do too.
Return to the Dunham Lab home page
Copyright 2004-2006 Maitreya Dunham
Last edit: 6 September 2006